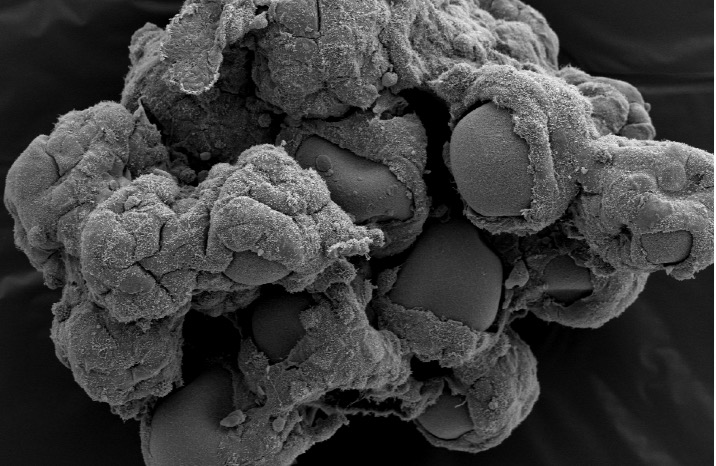
With threats like zika, ebola, SARS and tuberculosis increasing, researchers are using predictive tissue culture models to more accurately reflect how humans respond to pathogens.
The lack of sophisticated models has greatly slowed the ability to understand infectious disease from both the host and the pathogen perspective.
One approach to address this problem is the use of predictive tissue culture models. To develop these models, an understanding of how cells and tissues in our bodies function in a 3D context is required.
A research team from the Arizona State University Biodesign Institute team and NASA’s Johnson Space Center, have developed a realistic 3D co-culture model, which incorporates an important immune defense cell type found in the intestine, macrophages, key cells targeted by salmonella, a leading cause of food poisoning and systemic disease worldwide.
The researchers used strains of salmonella that cause gastroenteritis and life-threatening bloodstream infections, including the multidrug resistant salmonella strain D23580, which are responsible for devastating epidemics of invasive bloodstream infections in sub-Saharan Africa.
They used NASA bioreactor technology to develop 3D cell-based tissue models with increasing complexity to better recapitulate human tissues, bridging the gap between traditional cell culture and animal models currently used in infectious disease research.
This represents the first time that immune cells have been incorporated into a 3D intestinal model developed using the NASA rotating wall vessel bioreactor and the first application of this co-culture system to study infections that target the human gut.
“Our co-culture model thus offers a powerful new tool in understanding enteric [gut] pathogenesis and may lead to unexpected pathogenesis mechanisms and therapeutic targets that have been previously unobserved or unappreciated using other intestinal cell culture models,” Cheryl Nickerson, a researcher at ASU’s Biodesign Institute and professor in the School of Life Sciences, said in a statement.
Along with the 3D architecture and inclusion of immune cells, other complex factors of the tissue microenvironment must also be considered when building better tissue models for infections. These factors include physical forces, including the flow of fluids over the cell surface and different oxygen levels.
This allows researchers to capture the complex and dynamic interactions between the host and the pathogen, which govern the outcome of the infection process.
The researchers grew salmonella under different oxygen levels that it encounters before and during intestinal infection to better recapitulate the natural infection process. They also used dynamic bioreactor technology to culture cells under physiological fluid shear levels found in the intestinal tract.
“It’s a real challenge for researchers to accurately model all of the steps involved in the initiation and progression of host-pathogen interactions in the laboratory due to all of the complex factors in the human body that contribute to infection,” Jennifer Barrila, a lead co-author on the study, said in a statement.
“By harnessing the naturally low fluid shear environment generated by the NASA RWV bioreactor during model development combined with the use of physiologically relevant oxygen conditions during infection, we have come several steps closer to achieving our ultimate goal of recreating this complex 3D microenvironment.”
The response of the new model to infection with the different types of salmonella was very different for each strain, which demonstrates the model’s ability to distinguish between these closely related pathogens based on their infection characteristics.
Important differences were observed between the bacterial strains in model colonization and intracellular co-localization patterns in epithelial and immune cells.
“One important advantage of using this 3D multicellular in vitro host model system is the ability to visualize the co-localization patterns of different pathogens within the different host cell types,” Jiseon Yang, a lead co-author of the study, said in a statement. “I believe that these platforms can advance our knowledge of a variety of enteric diseases of both infectious (bacterial and viral) and non-infectious etiologies (IBS/IBD, drug toxicity, etc).”
Nickerson’s team is now focused on building in further complexity into the models so that layer by layer they can reach the ultimate goal of growing and fully mimicking whole 3D organs. They also want to add even more cells to the mix to provide researchers a hierarchical series of advanced new tools to study health and disease.
The study was published in npj Microgravity.




