
If you want to research historical events for a college essay, learn about tropical fish, or even translate text into a different language, you can type keywords into an internet search engine and get almost instant results drawn from diverse, international sources on that subject.
Unfortunately, it’s not so easy for the scientist trying to find solutions for the COVID-19 pandemic. Even though researchers across the world have already amassed a wealth of information about the disease and continue to reveal new insights every day, this valuable data is stored in different digital libraries, organized in different structures, and written with different jargon. To get the most out of our collective COVID-19 knowledge, someone needs to collect it all in one place.
And that’s precisely what a team led by Lawrence Berkeley National Laboratory (Berkeley Lab) is doing. Under a special project launched in May, computing and bioinformatics experts are working together to develop a platform that consolidates disparate COVID-19 data sources and uses the unified library to make predictions – about potential drug targets, for example.
“We’ve built what is called a knowledge graph, where we pull all the various heterogeneous kinds of biological data out there into one location, and organize it according to how the data relate to each other using techniques such as link prediction,” said project principal investigator Chris Mungall, head of the Biosystems Data Science department in Berkeley Lab’s Biosciences Area. “The next step, and what we are working on now, is to apply machine-learning approaches to our knowledge graph – named KG-COVID-19 – to predict what existing or new drugs could work against COVID-19 based on the properties of those compounds and the viral or human macromolecules they target.”
The knowledge graph is freely available on the project wiki and can be analyzed by free, open-source software. It currently includes data on approximately 32,000 drugs, 21,000 human and 272 viral proteins plus roughly the same number of genes, and more than 50,000 scientific studies and clinical trials. New and relevant information is added as it becomes available.
Rising to the challenge
Project member and technical lead Justin Reese explained that he and his colleagues were able to construct the COVID-19 knowledge graph in such a short time because they were already in the midst of developing the platform for a different project – involving drug prediction for cancer – when the pandemic hit.
“The crisis happened very quickly, but we knew from the outset that there was actually a fair bit of existing information about related coronaviruses that could be leveraged. And we knew that new research was going to kick into overdrive,” said Reese, a software developer in the Biosciences Area. “So, we thought, ‘OK, we know what to do; we’re good at creating knowledge graphs that make scattered data useful.’”
Reese, Mungall, and fellow team members Deepak Unni and Marcin Joachimiak set to work feeding the knowledge graph with datasets like those generated from genetic analyses, molecular structure models, and biochemical reaction studies. Then, they loaded in text-based knowledge that had been gathered by COVIDScholar, a natural language-processing machine-learning tool developed by another Berkeley Lab team.
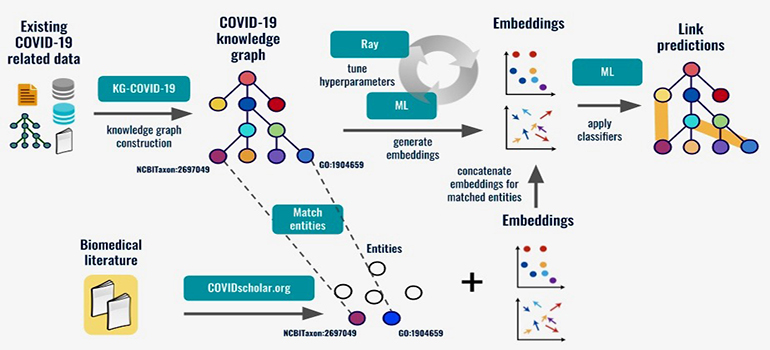
Workflow for using link prediction to extract actionable knowledge from the KG-COVID-19 knowledge graph. Data about COVID-19 is ingested to construct the knowledge graph, and machine learning algorithms are applied to predict new links, for example drugs that can be repurposed to treat COVID-19. (Credit: Berkeley Lab)
Once the knowledge graph was online, the scientists began using the high-performance computing resources at Berkeley Lab and Google Cloud to run predictive machine-learning algorithms. At the same time, they are working with external collaborators from academia and industry to develop a simple user interface, so that the knowledge graph can be used by doctors and medical researchers. As of now, the KG-COVID-19 is difficult to use without experience in bioinformatics.
The end goal is to create a resource that can make discoveries based on data connections that humans would take too long to find, and that can provide suggestions alongside search results. “One way you’ll be able to access our information is by asking our graph a question, like you would with Google, and getting results on your query and all related results along with context,” said Unni, a staff software developer. “So, if you’re trying to learn more about a particular viral protein, for example, alongside the direct results you’ll be presented with more information about molecules that might interact with that protein, and so on.”
The knowledge graph will be used to prioritize which potential drugs and drug targets are likely to be important, which could save significant time in the drug development process. This is done using machine-learning techniques that identify existing drugs that could be repurposed for COVID-19 treatment, and drug targets that are important in the disease process. Bench scientists and clinicians can then investigate these candidate drugs and drug targets for their usefulness in treating COVID-19.
According to the team, updated versions of the knowledge graph will be released monthly into the fall of 2020, but biomedical researchers are already using the first version. “The KG-COVID-19 project was launched less than three months ago but has already been leveraged to support DOE and international COVID-19 efforts,” said Reese. He noted that the tool is currently being integrated into the National Institute of Health’s National COVID Cohort Collaborative (N3C). “It’s gratifying to see that it’s already been helpful and that it will soon have even greater impact,” said Reese.
Other collaborators include Peter Robinson and Vida Ravanmehr at The Jackson Laboratory, Tiffany Callahan at the University of Colorado Denver, and Luca Cappelletti at the University of Milan. The KG-COVID-19 project is supported by the Laboratory Directed Research and Development (LDRD) Program and the U.S. Department of Energy National Virtual Biotechnology Laboratory: a consortium of DOE National laboratories with core capabilities relevant to the threats posed by COVID-19, and funded under the Coronavirus Aid, Relief, and Economic Security (CARES) Act.





Tell Us What You Think!