Imagine a robot that could help you tidy your home: roving about, sorting stray socks into the laundry and dirty dishes into the dishwasher. While such a practical helper may still be the stuff of science fiction, Caltech scientists have developed an autonomous molecular machine that can perform similar tasks — at the nanoscale. This “robot,” made of a single strand of DNA, can autonomously “walk” around a surface, pick up certain molecules and drop them off in designated locations.
The work was done in the laboratory of Lulu Qian, assistant professor of bioengineering. It appears in a paper in the Sept. 15 issue of Science.
“Just like electromechanical robots are sent off to faraway places, like Mars, we would like to send molecular robots to minuscule places where humans can’t go, such as the bloodstream,” says Qian. “Our goal was to design and build a molecular robot that could perform a sophisticated nanomechanical task: cargo sorting.”
Led by former graduate student Anupama Thubagere (PhD ’17), the researchers constructed three basic building blocks that could be used to assemble a DNA robot: a “leg” with two “feet” for walking, an “arm” and “hand” for picking up cargo, and a segment that can recognize a specific drop-off point and signal to the hand to release its cargo. Each of these components is made of just a few nucleotides within a single strand of DNA.
In principle, these modular building blocks could be assembled in many different ways to complete different tasks — a DNA robot with several hands and arms, for example, could be used to carry multiple molecules simultaneously.
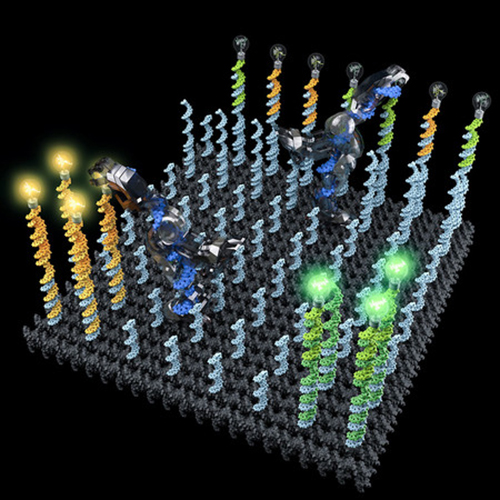

Conceptual illustration of two DNA robots collectively performing a cargo-sorting task on a DNA origami surface, transporting fluorescent molecules with different colors from initially unordered locations to separated destinations. Considerable artistic license has been taken. Image: Demin Liu
In the work described in the Science paper, the Qian group built a robot that could explore a molecular surface, pick up two different molecules — a fluorescent yellow dye and a fluorescent pink dye — and then distribute them to two distinct regions on the surface. Using fluorescent molecules enabled the researchers to see if the molecules ended up in their intended locations. The robot successfully sorted six scattered molecules, three pink and three yellow, into their correct places in 24 hours. Adding more robots to the surface shortened the time it took to complete the task.
“Though we demonstrated a robot for this specific task, the same system design can be generalized to work with dozens of types of cargos at any arbitrary initial location on the surface,” says Thubagere. “One could also have multiple robots performing diverse sorting tasks in parallel.”
The key to designing DNA machines is the fact that DNA has unique chemical and physical properties that are known and programmable. A single strand of DNA is made up of four different molecules called nucleotides — abbreviated A, G, C, and T — and arranged in a string called a sequence. These nucleotides bond in specific pairs: A with T, and G with C. When a single strand encounters a so-called reverse complementary strand — for example, CGATT and AATCG — the two strands zip together in the classic double helix shape.
A single strand containing the right nucleotides can force two partially zipped strands to unzip from each other. How quickly each zipping and unzipping event happens and how much energy it consumes can be estimated for any given DNA sequence, allowing researchers to control how fast the robot moves and how much energy it uses to perform a task. Additionally, the length of a single strand or two zipped strands can be calculated. Thus, the leg and foot of a DNA robot can be designed for a desired step size — in this case, 6 nanometers, which is about a hundred millionth of a human’s step size.
Using these chemical and physical principles, researchers can design not only robots but also “playgrounds,” such as molecular pegboards, to test them on. In the current work, the DNA robot moves around on a 58-nanometer-by-58-nanometer pegboard on which the pegs are made of single strands of DNA complementary to the robot’s leg and foot. The robot binds to a peg with its leg and one of its feet — the other foot floats freely. When random molecular fluctuations cause this free foot to encounter a nearby peg, it pulls the robot to the new peg and its other foot is freed. This process continues with the robot moving in a random direction at each step.
It may take a day for a robot to explore the entire board. Along the way, as the robot encounters cargo molecules tethered to pegs, it grabs them with its “hand” components and carries them around until it detects the signal of the drop-off point. The process is slow, but it allows for a very simple robot design that utilizes very little chemical energy.
“We don’t develop DNA robots for any specific applications. Our lab focuses on discovering the engineering principles that enable the development of general-purpose DNA robots,” says Qian. “However, it is my hope that other researchers could use these principles for exciting applications, such as using a DNA robot for synthesizing a therapeutic chemical from its constituent parts in an artificial molecular factory, delivering a drug only when a specific signal is given in bloodstreams or cells, or sorting molecular components in trash for recycling.”
The paper is titled “A cargo-sorting DNA robot.” In addition to Thubagere and Qian, other co-authors are postdoctoral scholar Wei Li, graduate student Robert Johnson (BS ’15), former visiting student Zibo Chen, former undergraduates Shayan Doroudi (BS ’13), Yae Lim Lee (BS ’12), Gregory Izatt (BS ’14), and Sarah Wittman (BS ’13); former graduate student Niranjan Srinivas (PhD ’15), former senior research fellow Damien Woods; and Erik Winfree (PhD ’98), professor of computer science, computation and neural systems, and bioengineering. Funding was provided by Caltech Summer Undergraduate Research Fellowships, the National Science Foundation, and the Burroughs Wellcome Fund.
Source: Caltech