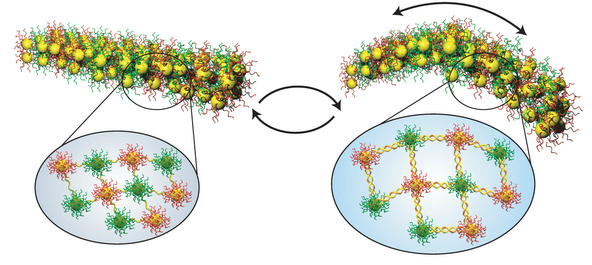
The study’s nanoscale muscles are comprised of gold nanoparticles, which are connected by single-stranded DNA.
The base pairs found in DNA are key to its ability to store protein-coding information, but they also give the molecule useful structural properties. Getting two complementary strands of DNA to zip up into a double helix can serve as the basis of intricate physical mechanisms that can push and pull molecular-scale devices.
Engineers at the University of Pennsylvania have developed nanoscale “muscles” that work on this principle. By carefully incorporating strands of custom DNA into different layers of flexible films, they can force those films to bend, curl and even flip over by introducing the right DNA cue. They could also reverse these changes by way of different DNA cues.
One day, the flexing of these muscles could be used in diagnostic devices, capable of signaling changes in gene expression from within cells.
The researchers demonstrated this system in a study published in Nature Nanotechnology.
The study was led John C. Crocker and Daeyeon Lee, professors of Chemical and Biomolecular Engineering in Penn’s School of Engineering and Applied Science, along with Tae Soup Shim, who was then a post-doctoral associate in both researchers’ groups. David Chenoweth, an assistant professor of chemistry in Penn’s School of Arts & Sciences, and So-Jung Park, a professor in the Department of Chemistry and Nano Science at Ewha Womans University, Seoul, also contributed to the study. Other Penn co-authors include Zaki G. Estephan, Zhaoxia Qian, Jacob H. Prosser and Su Yeon Lee, graduate students and postdoctoral researchers in the departments of Chemical and Biomolecular Engineering, Materials Science and Engineering and Chemistry.
The nanoscale muscles in the study are comprised of gold nanoparticles, which are connected to one another by single-stranded DNA. The researchers built up the films layer-by-layer, introducing different sets of DNA-linked nanoparticles at different depths. Each set of nanoparticles contained links with different sequences.
“The way the actuation works,” Crocker says, “is that we add single-stranded DNA that is complementary to a portion of the bridges between the particles. When that DNA diffuses in, it turns just those bridges into double-stranded DNA helices.”
Because the specific sequence of the added DNA is tailored to match different sets of nanoparticle bridges, the researchers could target individual layers of the film, forming double-stranded bridges in just those layers.
This mechanism was critical to getting the films to flex, as single-stranded and double-stranded bridges are different lengths.
“It so happens that double-stranded DNAs are longer than single-stranded DNAs with the same number of bases,” Crocker says, “so when the added strand binds, the bridge gets a little longer and the material expands. If only one layer of the film expands, the film curls.”
The researchers also designed a way of reverting the bridges to their original, single-stranded state, undoing this curl. The strands that give the curling cue also have a “handle” that does not bind to the bridges. Pulling on this handle splits apart the double helix the added DNA forms.
“We make the strand that we added to expand the bridges a little longer than it needs to be,” Crocker says. “After it forms a double helix with the bridge, there are another 7 bases of leftover single-stranded DNA dangling off to the side of the bridge. To reverse the process, we add a ‘stripper’ strand that is complementary to the ‘expander’ strand and the extra dangling ‘handle.’ It actually hybridizes to the dangling handle, and then pulls the expander strand off the bridge, forming a double helix in solution that floats away, allowing the bridge to revert to its shorter, single-stranded form.”
Getting the films to curl or to flip over entirely is just a proof-of-concept for now, but this muscle-like flexing behavior could have a host of applications at the nanoscale.
Being able to respond to one cue and totally ignore the other — impossible for systems that flex based on temperature or acidity changes — is critical for their ability to work as diagnostic devices.
“A ‘far-out’ application we have thought of is in intracellular situations where we can’t exactly control things with wires or wireless equipment, Crocker says. “We could make a device that absorbs or reflects a certain wavelength of light based upon the spacing of its internal layers, and we could then alter that spacing using a chemical signal. This signal could be a messenger RNA, so the device provides a single-cell gene expression readout. These intracellular devices could be read out under a microscope, or inside the body using infrared imaging.”
The research was supported by the National Science Foundation under an MRSEC seed award DMR11-20901 and the National Research Foundation of Korea, under grants NRF-2015R1A2A2A01003528 and 2016R1C1B2016089.
Shim is now an assistant professor in the Department of Chemical Engineering at Ajou University, Suwon, Korea.
Source: UPenn




