By Marilyn Matz, CEO and Co-Founder, and Zachary Pitluk Ph.D., V.P. of Life Sciences and Healthcare
Paradigm4

Marilyn Matz
Precision medicine is poised to transform healthcare, with biobanks being identified as a key enabler to bring it to fruition by 20301 by putting new, extended datasets at the disposal of pharmaceutical researchers and healthcare professionals. Organizations are working to facilitate more collaboration when working with the vast data stored in biobanks, with the International Hundred Thousand Plus Cohort Consortium (IHCC) bringing together over 100 cohorts in 43 countries comprising over 50 million participants. This is almost two orders of magnitude larger than the biggest single cohort today2 and could have an exponential impact on global scientific research.
In order to realize the potential of biobank data to transform drug R&D and precision medicine, rapid, cost-effective and scalable data analytics and management approaches are becoming essential. Current data storage, analytics platforms and computing approaches overlook too much of the data wrangling and distributed computing tasks while also incurring performance bottlenecks. Here, we look at how disruptive technologies offer a fundamentally new approach to addressing the challenges of population-scale data management and reducing the cost of biobank data analysis.

Zachary Pitluk Ph.D.
Investigating genetic predisposition
Biobanks are ground-breaking resources that have evolved considerably since their inception three decades ago. The data stored and shared, from simple health records and lifestyle information in the early years, to the unprecedented levels of genetic information available today, is used in many areas of public health, such as drug and
biomarker development, clinical trials, monitoring of the effects of drugs on eventual outcomes, and clinical care for therapy. Much of the progress biobanks have facilitated in precision medicine so far has happened following the rise of genomics and the ability to develop large, electronic databases that store vast amounts of information associated with patient samples.3
Many countries either have or are developing national biobanks as part of long-term studies to investigate the contributions of genetic predisposition and environmental exposure to the development of disease. This includes the UK (UK Biobank), China (Kadorrie), Japan (Jenger), the US (All of Us) and Finland (FinnGen). The UK biobank (UKBB) has 7,400 categories of phenotypes along with single nucleotide polymorphisms (SNP) and whole exome sequencing (WES) data from 500,000 participants – an important milestone in the availability of population health data. It is now embarking on whole genome sequencing and proteomics.
National biobanks have also played a part in COVID-19 research, with UKBB data being used in a COVID-19 antibody study to determine the extent of past infection rates across the UK.4 The study found that 99% of participants who had tested positive for previous infection retained antibodies to SARS-CoV-2 for 3 months after being infected – an early indication that any vaccine produced that stimulated antibody production was likely to offer protection against infection. The UKBB is now embarking on another study to determine the long-term health impacts of COVID-19. Although it is a respiratory disease, evidence is showing that it can affect other parts of the body, including the heart, liver, kidneys and brain. Researchers from UKBB are investigating the direct effects of the infection on changes in the structure and function of these organs using data from 1,500 people.5
Building a biobank
The real value of a biobank lies in two key areas: using genetic evidence to uncover new targets and biomarkers and broadening the scope of a drug under development to potentially expand and find new uses for drugs that are already on the market. Typically, genetic information such as WGS (whole genome sequencing), WES and SNP (Single nucleotide polymorphisms) is aggregated with a range of other data on the same individuals: health records like GP data, hospitalizations, diagnoses, prescriptions, MRI (magnetic resonance imaging), lab results from biochemistry and hematology, and patient-reported data such as family history, behavioral history, and socio-demographics. These data are used for computing large association analyses that associate specific genetic variations and specific phenotypes with susceptibility to or protection from certain diseases. This helps to identify genetic markers that are candidates for drug targets and biomarkers that can be used to predict the presence or future development of a disease.
Discovering new uses for existing drugs – known as drug repositioning or indication expansion – offers substantial advantages in terms of cost, time and risk, and represents a highly attractive strategy for drug discovery. In the case of rare diseases, whose pathophysiology is often poorly characterized, computational techniques for predictive expansion offer a quick way of identifying testable hypotheses that may be translated into the clinic. Large-scale genome-sequencing initiatives contribute to identifying the genetic variation/s responsible for the disorder, and opening opportunities to rapidly repurpose drugs that target the correspondent protein/s.6
The information generated is made available to medical research projects and, depending on the project, different biobanks have different guidelines and governance for using the data. However, organizations using biobanks must adhere to the appropriate regulatory and ethical standards when they access data.
Data wrangling
While the expanding potential of biobanks brings opportunities, the data they compile is increasingly complex. Considering biobanks store data from hundreds or thousands of individuals, scientists require the computation power to handle billions of data points. But it is not just a numbers game. Researchers are seeking connections across datasets, examining the information, asking questions of the data to test and validate a hypothesis, or looking for a key to unlock a new development. This is the fundamental challenge in data science – how to best marry computer science expertise with biological insights.7
There is a significant risk that researchers spend too much time navigating complex data management and optimizing scalable computing to the detriment of their scientific output. Many genomic data analysis methods, while computationally demanding, are ‘data parallel’ in nature. ‘Data parallel’ refers to applications that can be divided into independent tasks which can be distributed across distinct computational nodes8, for example, in linkage disequilibrium analysis and GWAS. Performing GWAS and phenome-wide association studies (PheWAS) to identify potential connections between genetic variants and phenotypes is an increasingly computationally challenging task where a data analytics engine that offers workflow or dataflow optimization without requiring user involvement, would be a valuable option.
Most organizations struggle to provide their scientists with a unified platform for systemic analysis of biobank data at multiple levels simultaneously, including the genome, phenome, transcriptome and epigenome, in a way that balances respecting patient privacy and sharing data appropriately with other collaborators. Traditional database management systems, distributed computing systems, data lakes and file-based systems all exhibit several constraints that limit the value to be gained from genomic biobank analysis. (Table 1) For example, they are often too slow to process heterogeneous scientific datasets, and as systems grow in complexity, databases can easily become isolated and data sharing between systems is restricted.
As the volume of data being collected in biobanks continues to increase, the magnitude of the corresponding computing complexity increases polynomially. Today, a single powerful server could easily spend weeks computing the results of one large-scale GWAS or PheWAS study. With such rich and vast source of information available, researchers require a solution that can reduce the time-to-results by orders of magnitude and provide the agility to allow many computations to be completed interactively within a single workday.
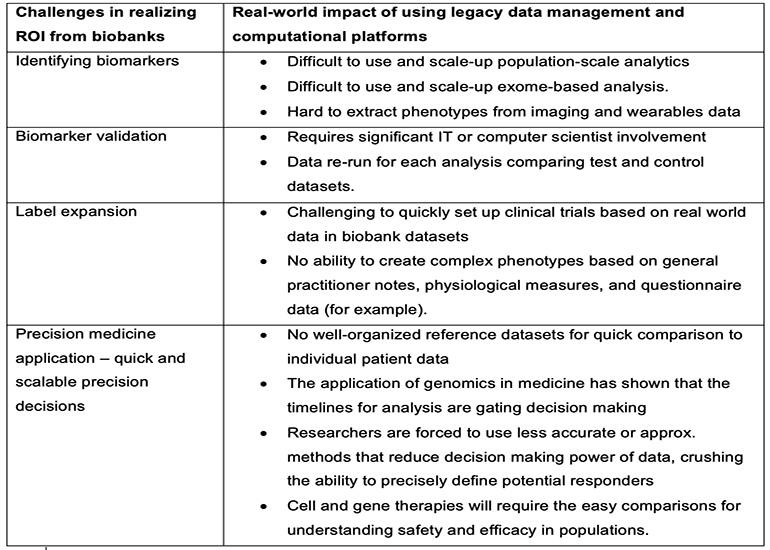
Table 1 summarizes some of the challenges in using legacy data management and computational platforms for genomic biobank analysis.
Case study: The Global Biobank Engine
The Global Biobank Engine (GBE)9 uses the SciDB scientific computing platform with REVEAL Biobank application (Paradigm4) for large-scale joint analysis of large genotype-phenotype datasets like the UKBB and has shown superior performance with the large amount of data that needs to be stored and queried.10
Funded by Stanford University, the GBE was built using a modified version of the GBE ExAC browser code-base and an extended version of the IDB Exomes Browser. The GBE development was led by Prof. Manuel Rivas, and the engine supports browsing for results from GWAS and PheWAS, gene-based tests, and genetic correlation between phenotypes.
The aim of the GBE is to provide a platform that facilitates the dissemination of summary statistics from biobanks to the scientific and clinical communities. Implementing SciDB has facilitated the analysis for an array of research projects whose results are displayed in the GBE, including:
- Testing for associations between PTVs and 135 different medical phenotypes including cancers and complex diseases among 337,205 participants in the UKBB.11
- Identifying key latent components of genetic associations and the contributions of variants, genes, and phenotypes to each component, by applying truncated singular value decomposition (DeGAs) to matrices of summary statistics derived from genome-wide association analyses across 2,138 phenotypes measured in 337,199 white British individuals in a UKBB study.12
- Presenting a method for identifying associations between genetic variation and disease that considers multiple rare variants and phenotypes at the same time, therefore improving the ability to identify rare variants associated with disease compared to considering a single rare variant and a single phenotype.13
Meeting the needs of biobank users
The rapid pace of innovation in multi-omics, coupled with the decreasing cost of some technologies, indicates a change in the way we understand disease and deliver optimized treatment. As the storage and use of genetic and phenotypic data scales up, the time and effort it takes to manage, analyze, and run advanced AI algorithms cannot increase with it. The biobank community needs a cost-efficient, fast platform that can handle the complex, heterogeneous data from proprietary and public datasets.
Users looking to leverage large-scale genomic data, such as pharmaceutical companies for drug discovery or indication expansion, or healthcare organizations gearing towards precision medicine, can benefit from implementing an end-to-end, application specific computational solution. The growing research results of large pharmaceutical companies (such as Alnylam Pharmaceuticals, which published six posters and research papers using biobank data in less than one year), as well as biotech start-ups, demonstrates how emerging technologies can transform the pace and capabilities of daily research.
Indeed, this collaborative landscape being fostered by genetics companies, pharmaceutical companies, healthcare organizations, and data analytics experts will be vital to the future success of genomic biobank initiatives.
For more information, visit paradigm4.com or contact [email protected].
References:
- Denny JC, Collins FS. Precision medicine in 2030—seven ways to transform healthcare. Cell. 2021 DOI: doi.org/10.1016/j.cell.2021.01.015
- Manolio TA, Goodhand P, Ginsburg G. The International Hundred Thousand Plus Cohort Consortium: integrating large-scale cohorts to address global scientific challenges. Lancet Digit Health. 2020; 2: e567-e568
- Coppola L, Cianflone A, Grimaldi AM, et al. Biobanking in health care: evolution and future directions. J Transl Med. 2019; 17(1):172.
- UK Biobank COVID-19 antibody study: final results, uk (2021) gov.uk/government/statistics/uk-biobank-covid-19-antibody-study-final-results/uk-biobank-covid-19-antibody-study-final-results
- Covid: UK Biobank scans aim to reveal health legacy, BBC (11 March 2021) bbc.co.uk/news/health-56352138
- Pushpakom S, Iorio F, Eyers PA, et al. Drug repurposing: progress, challenges and recommendations. Nat Rev Drug Discov. 2019;18:41–58.
- Dash S, Shakyawar SK, Sharma M, et al. Big data in healthcare: management, analysis and future prospects. J Big Data, 2019; 6(54).
- Sikorska K, Lesaffre E, Groenen PF, Eilers PH. GWAS on your notebook: fast semiparallel linear and logistic regression for genome-wide association studies. BMC Bioinformatics. 2013; 14:166.
- Global Biobank Engine, Stanford University. biobankengine.stanford.edu/
- McInnes G, Tanigawa Y, DeBoever C, et al. Global Biobank Engine: enabling genotype-phenotype browsing for biobank summary statistics, Bioinformatics, 2019; 35(14):2495–2497.
- DeBoever C, Tanigawa Y, Lindholm ME, et al. Medical relevance of protein-truncating variants across 337,205 individuals in the UK Biobank study. Nat Commun 9, 2018;1612.
- Tanigawa Y, Li J, Justesen JM, et al. Components of genetic associations across 2,138 phenotypes in the UK Biobank highlight adipocyte biology. Nat Commun 10, 2019;4064.
- DeBoever C, Aguirre M, Tanigawa Y, et al. Bayesian model comparison for rare variant association studies of multiple phenotypes, bioRxiv 257162; 2018.





Tell Us What You Think!