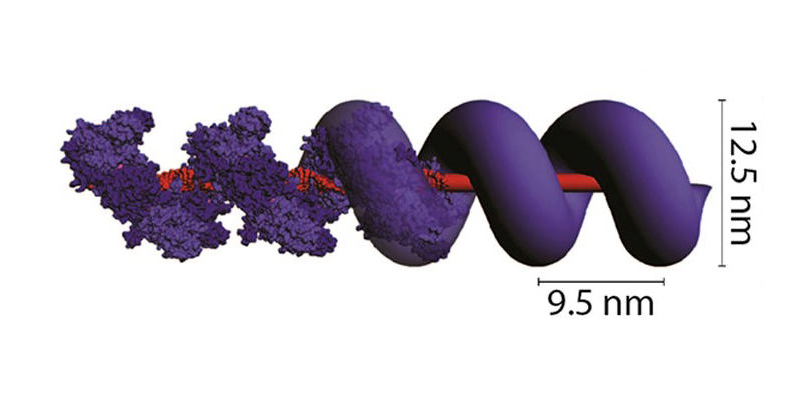
The protein RecA (purple units), wraps around and fortifies double-stranded DNA, enabling scientists to build large structures with the genetic material. Credit: NIST
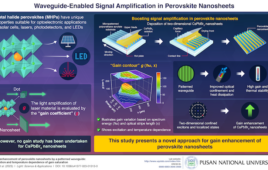
DNA is the stuff of life, but it is also the stuff of nanotechnology. Because molecules of DNA with complementary chemical structures recognize and bind to one another, strands of DNA can fit together like Lego blocks to make nanoscale objects of complex shape and structure.
But researchers need to work with much larger assemblages of DNA to realize a key goal: building durable miniature devices such as biosensors and drug-delivery containers. That’s been difficult because long chains of DNA are floppy and the standard method of assembling long chains is prone to error.
Using a DNA-binding protein called RecA as a kind of nanoscale rebar, or reinforcing bar, to support the floppy DNA scaffolding, researchers at the National Institute of Standards and Technology (NIST) have constructed several of the largest rectangular, linear and other shapes ever assembled from DNA. The structures can be two to three times larger than those built using standard DNA self-assembly techniques.
In addition, because the new method requires fewer chemically distinct pieces to build organized structures than the standard technique, known as DNA origami, it is likely to reduce the number of errors in constructing the shapes. That’s a big plus for the effort to produce reliable DNA-based devices in large quantities, said NIST researcher Alex Liddle.
Although RecA’s ability to bind to double-stranded DNA has been known for years, the NIST team is the first to integrate filaments of this protein into the assembly of DNA structures. The addition of RecA offers a particular advantage: Once one unit of the protein binds to a small segment of double-stranded DNA, it automatically attracts other units to line up alongside it, in the same way that bar magnets will join end-to-end. Like bricks filling out a foundation, RecA lines the entire length of the DNA strand, stretching, widening and strengthening it. A floppy, 2-nanometer-wide strand of DNA can transform into a rigid structure more than four times as wide.
“The RecA method greatly extends the ability of DNA self-assembly methods to build larger and more sophisticated structures,” said NIST’s Daniel Schiffels.
Schiffels, Liddle and their colleague Veronika Szalai describe their work in a recent article in ACS NANO.
The new method incorporates the DNA origami technique and goes beyond it, according to Liddle. In DNA origami, short strands of DNA that have a specific sequence of four base pairs are used as staples to tie together long sections of DNA. To make the skinny DNA skeleton stronger and thicker, the strand may loop back on itself, quickly using up the long string.
If DNA origami is all about the folding, Liddle likened his team’s new method to building a room, starting with a floor plan. The location of the short, single-stranded pieces of DNA that act as staples mark the corners of the room. Between the corners lies a long, skinny piece of single-stranded DNA. The enzyme DNA polymerase transforms a section of the long piece of single-stranded DNA into the double-stranded version of the molecule, a necessary step because RecA only binds strongly to double-stranded DNA. Then RecA assembles all along the double strand, reinforcing the DNA structure and limiting the need for extra staples to maintain its shape.
With fewer staples required, the RecA method is likely able to build organized structures with fewer errors than DNA origami, Liddle said.
The research was a collaboration between NIST’s Center for Nanoscale Science and Technology and the Maryland NanoCenter at the University of Maryland at College Park.
SOURCE: NIST