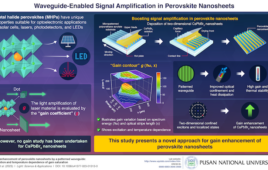
A cluster of microsponges made of long strands of folded RNA, as seen by scanning electron microscopy. Image: Hammond laboratory |
For
the past decade, scientists have been pursuing cancer treatments based
on RNA interference—a phenomenon that offers a way to shut off
malfunctioning genes with short snippets of RNA. However, one huge
challenge remains: finding a way to efficiently deliver the RNA.
Most
of the time, short interfering RNA (siRNA)—the type used for RNA
interference—is quickly broken down inside the body by enzymes that
defend against infection by RNA viruses.
“It’s
been a real struggle to try to design a delivery system that allows us
to administer siRNA, especially if you want to target it to a specific
part of the body,” says Paula Hammond, the David H. Koch Professor in
Engineering at MIT.
Hammond
and her colleagues have now come up with a novel delivery vehicle in
which RNA is packed into microspheres so dense that they withstand
degradation until they reach their destinations. The new system,
described Feb. 26 in the journal Nature Materials,
knocks down expression of specific genes as effectively as existing
delivery methods, but with a much smaller dose of particles.
Such
particles could offer a new way to treat not only cancer, but also any
other chronic disease caused by a “misbehaving gene,” says Hammond, who
is also a member of MIT’s David H. Koch Institute for Integrative Cancer
Research. “RNA interference holds a huge amount of promise for a number
of disorders, one of which is cancer, but also neurological disorders
and immune disorders,” she says.
Lead
author of the paper is Jong Bum Lee, a former postdoc in Hammond’s lab.
Postdoc Jinkee Hong, Daniel Bonner PhD ’12 and Zhiyong Poon PhD ’11 are
also authors of the paper.
Genetic disruption
RNA
interference is a naturally occurring process, discovered in 1998, that
allows cells to fine-tune their genetic expression. Genetic information
is normally carried from DNA in the nucleus to ribosomes, cellular
structures where proteins are made. siRNA binds to the messenger RNA
that carries this genetic information, destroying instructions before
they reach the ribosome.
Scientists
are working on many ways to artificially replicate this process to
target specific genes, including packaging siRNA into nanoparticles made
of lipids or inorganic materials such as gold. Though many of those
have shown some success, one drawback is that it’s difficult to load
large amounts of siRNA onto those carriers, because the short strands do
not pack tightly.
To
overcome this, Hammond’s team decided to package the RNA as one long
strand that would fold into a tiny, compact sphere. The researchers used
an RNA synthesis method known as rolling circle transcription to
produce extremely long strands of RNA made up of a repeating sequence of
21 nucleotides. Those segments are separated by a shorter stretch that
is recognized by the enzyme Dicer, which chops RNA wherever it
encounters that sequence.
As
the RNA strand is synthesized, it folds into sheets that then
self-assemble into a very dense, sponge-like sphere. Up to half a
million copies of the same RNA sequence can be packed into a sphere with
a diameter of just two microns. Once the spheres form, the researchers
wrap them in a layer of positively charged polymer, which induces the
spheres to pack even more tightly (down to a 200-nm diameter) and
also helps them to enter cells.
After
the spheres enter a cell, the Dicer enzyme chops the RNA at specific
locations, releasing the 21-nucleotide siRNA sequences.
Peixuan
Guo, director of the NIH Nanomedicine Development Center at the
University of Kentucky, says the most exciting aspect of the work is the
development of a new self-assembly method for RNA particles. Guo, who
was not part of the research team, adds that the particles might be more
effective at entering cells if they were shrunk to an even smaller
size, closer to 50 nm.
Targeting tumors
In the Nature Materials
paper, the researchers tested their spheres by programming them to
deliver RNA sequences that shut off a gene that causes tumor cells to
glow in mice. They found that they could achieve the same level of gene
knockdown as conventional nanoparticle delivery, but with about
one-thousandth as many particles.
The
microsponges accumulate at tumor sites through a phenomenon often used
to deliver nanoparticles: The blood vessels surrounding tumors are
“leaky,” meaning that they have tiny pores through which very small
particles can squeeze.
In
future studies, the researchers plan to design microspheres coated with
polymers that specifically target tumor cells or other diseased cells.
They are also working on spheres that carry DNA, for potential use in
gene therapy.
Self-assembled RNA interference microsponges for efficient siRNA delivery