People could start receiving life-saving treatment for cancer and other diseases much earlier with the help of a new method to analyze blood proteins.
A paper by researchers at Pacific Northwest National Laboratory describes a new way to prepare and analyze blood samples that significantly improves researchers’ ability to detect proteins that exist in very low concentrations — as little as 10 picograms per milliliter of blood. Such tiny, hard-to-detect concentrations are common with disease-indicating proteins, which are also called biomarkers.
Many researchers identify proteins by using antibodies, Y-shaped proteins that grab onto proteins they’re specifically designed to target. But many proteins either don’t have matching antibodies or the antibodies they do have aren’t good enough to find proteins when few of them are present.
Researchers have long used antibodies this way, but mass spectrometry provides an attractive alternative that is more effective at identifying proteins. To analyze a given protein in blood or other biological samples, researchers use a mass spectrometry approach called selected reaction monitoring, also known as SRM. SRM specifically looks for unique peptides that indicate the presence of a targeted protein.
In 2012, PNNL scientists improved upon standard SRM when they developed a new method called PRISM-SRM, which separates a sample into 96 fractions and determines which of those fractions contains targeted proteins. This enables researchers to run an enriched sample with more targeted proteins through the mass spectrometer. As a result, it can detect proteins that exist in concentrations between 1 nanogram and 100 picograms per milliliter in blood. The 100 picogram-level of sensitivity occurs when common proteins that aren’t related to disease are removed through a process called depletion, while 1 nanogram-level sensitivity occurs when common proteins aren’t depleted.
But they still wanted to detect proteins that are present in even smaller amounts. To run a sample with even more of the desired proteins through a mass spectrometer, a team of PNNL researchers led by Tao Liu created a new process that starts with larger sample quantities upfront. After cutting the initial sample into 96 fractions, the scientists isolated the fraction with the targeted protein similar to as they had done with PRISM. Next, they separated just that fraction into another 96 – and again isolated the fraction containing the protein of interest.
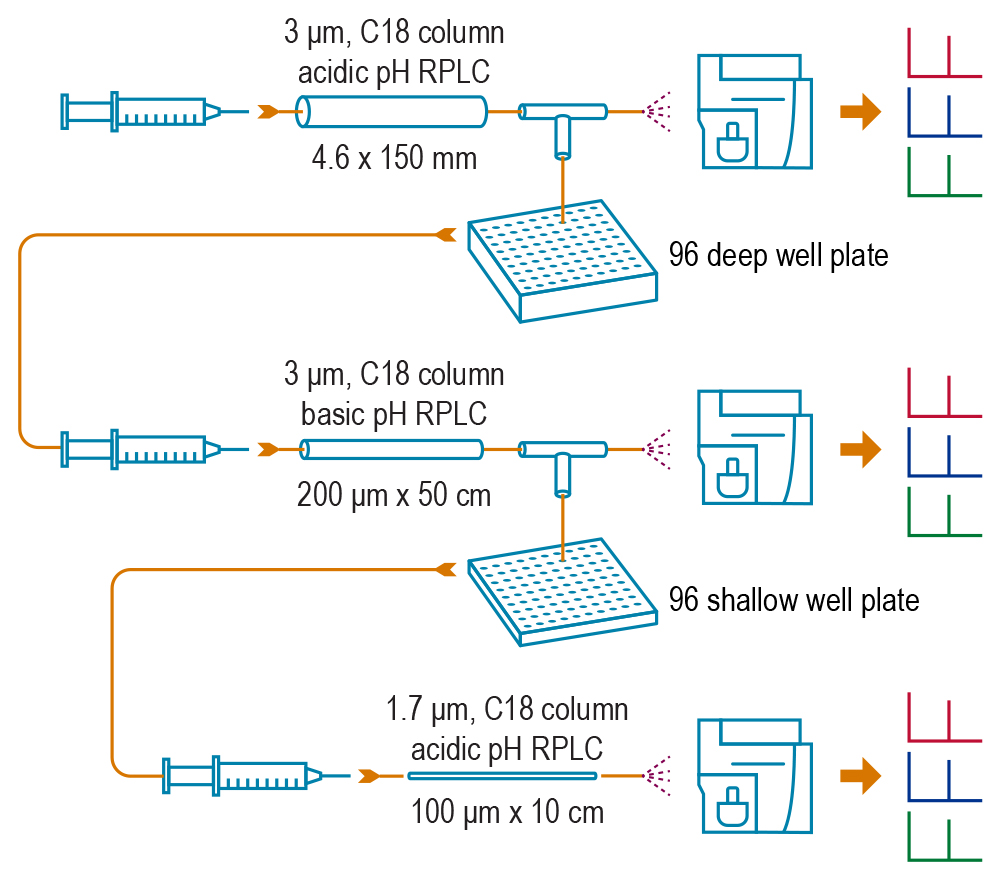
PNNL’s deep-dive SRM method offers a new way to prepare and analyze blood samples, and makes it possible to detect proteins that exist in very low concentrations – as little as 10 picograms per milliliter of a blood sample. The method has a three-step process: 1) Cut the initial sample into 96 fractions and isolate the fraction with the protein of interest, 2) Separate just that fraction into another 96 fractions and again isolate the fraction containing the protein of interest, and 3) Inject the resulting enriched sample into a mass spectrometer to analyze the proteins present. Image: Pacific Northwest National Laboratory
Called deep-dive SRM, this new approach makes it possible to target proteins that are present in the original blood sample in concentrations as little as 10 picograms per milliliter — and without needing antibodies or upfront depletion. That increases this method’s sensitivity by about 100,000 times when compared to conventional SRM analysis and by about 100 times when compared to PRISM-SRM.
Deep-dive SRM could be used by researchers to identify disease-indicating proteins in any biological or clinical sample before they become more plentiful — including potentially diagnosing cancer before a tumor is detectable by other means. Such early detection can increase the chances of saving a patient’s life.




