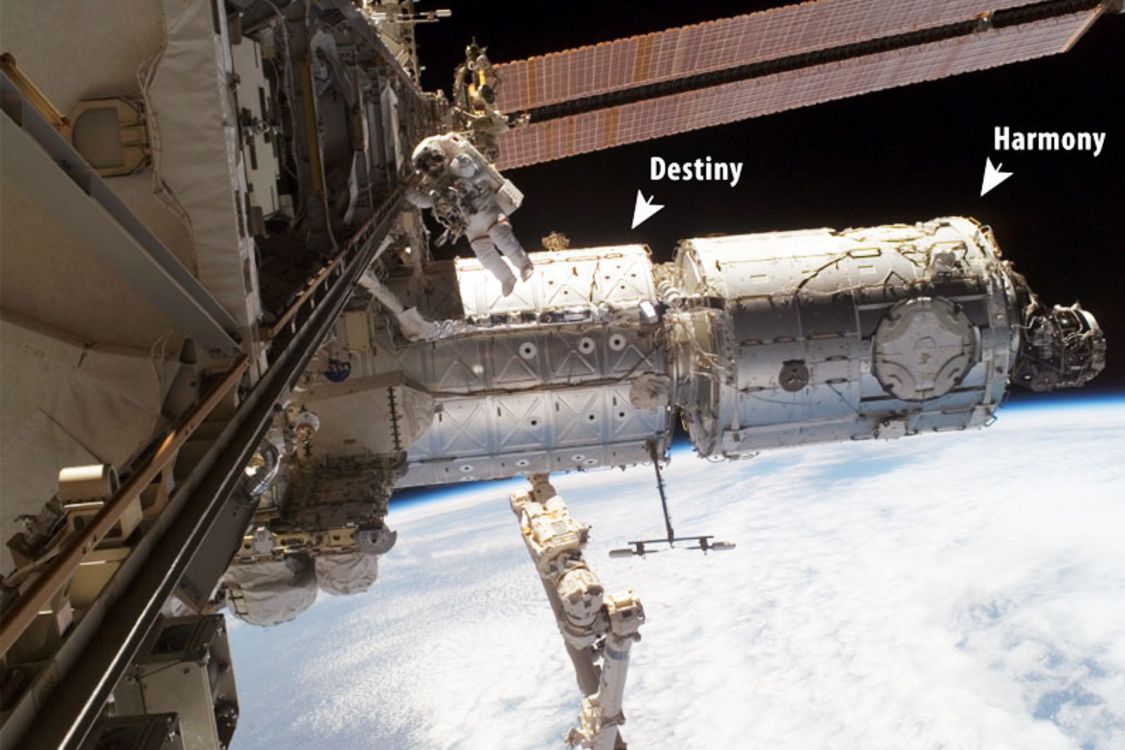
NASA astronaut Peggy Whitson walks in space outside the Destiny and Harmony modules of the International Space Station. Credit: Canadian Space Agency
Scientists are working to better understand how—and which—bacteria thrive aboard the International Space Station (ISS).
Researchers from the University of Montreal and McGill University created a new genomic methodology that can analyze how a bacterial ecosystem works under the harsh environments aboard the ISS. The novel method revealed a complex bacterial ecosystem at work on the ISS.
Until now, relatively little was known about the different types of microbes found on the space station. Researchers previously analyzed 16S ribosomal RNA (rRNA) gene amplification data for microbial barcoding. However, this type of analysis can be inaccurate across complex environmental samples due to the intra-genomic variation that is often present.
To overcome these challenges, the researchers developed a new approach dubbed ANCHOR, a 16S rRNA gene amplicon pipeline for microbial analysis of multiple environmental sample. This new technique—which is designed specifically for improved species-level microbial identification—uses paired-end sequences directly, multiple high-complexity samples and multiple reference databases.
“ANCHOR is designed with utility for non-ideal, uncharacterized biology in mind and, in particular, to provide flexibility to complex data sets and complement high uncertainty meta-transcriptomics,” the team wrote in the study.
The researchers used this multipronged method to directly test a real-world data set from surface swabs of ISS and identified 714 putative unique microbiome species or strains. They also performed differential abundance analysis and found that the astronaut sleeping quarters were dominated by human gastrointestinal tract bacteria, while the laboratory space bacteria derived from non-human animals present on the shuttle.
By taking a new approach, the researchers are hoping to identify and map different species inside of the space station, with the goal of safeguarding the astronaut’s health, as well as take future precautions for long-term space travel.
NASA has been attempting to reduce the amount of microbial growth on ISS since it was first launched into space in 1998. There has been strict clean and decontamination protocols in place such as regular cleanings and vacuuming of the space station’s living and working quarters.
However, when resupply missions arrive with food supplies, lab equipment and live plants and animals, new bacteria species are continually being added. The new bacteria, combined with existing bacteria aboard the space station, build-up and can cause significant health issues for the astronauts.
“Scientists have a well-documented understanding of broad bacterial families on the ISS, but now we’ve discovered a more diverse bacterial ecosystem that we ever expected,” Emmanuel Gonzalez, a metagenomic specialist at McGill, said in a statement. “It’s an exciting step forward in understanding the biosphere that will accompany humans into extra-terrestrial habitats.”The team believes the research will also have applications for environmental management and healthcare. If researchers are able to replicate this approach on Earth, they could address many environmental challenges in the oceans and soils, and apply better techniques for addressing human diseases and microbiomes.
“The new methodology provides us spectacular snapshots of the bacterial world in space and the possibilities of applying this method to explore new microbiome environments are really exciting,” Nicholas Brereton, a researcher at UdeM’s Institut de recherche en biologie végétale, said in a statement.
The study was published in Environmental Microbiology.