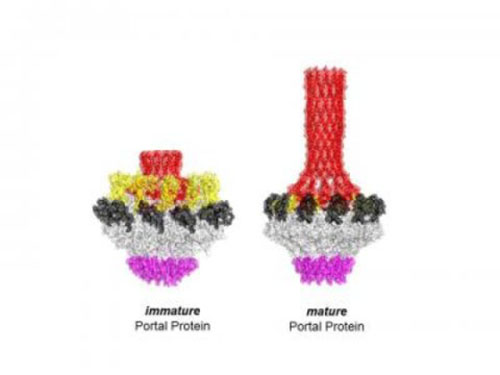
Image: Cingolani Lab
To generate swarms of new viral particles, a virus hijacks a cell into producing masses of self-assembling cages that are then loaded with the genetic blueprint for the next infection. But the picture of how that DNA is loaded into those viral cages, or capsids, was blurry, especially for two of the most common types of DNA virus on earth, bacterial viruses and human herpesvirus. Thomas Jefferson University researchers pieced together the three-dimensional atomic structure of a doughnut-shaped protein that acts like a door or “portal” for the DNA to get in and out of the capsid, and have now discovered that this protein begins to transform its structure when it comes into contact with DNA. Their work has been published in Nature Communications.
“Researchers thought that the portal protein acts as an inert passageway for DNA,” says senior author Gino Cingolani, Ph.D., a Professor in the Department of Biochemistry and Molecular Biology at Thomas Jefferson University and researcher at the Sidney Kimmel Cancer Center. “We have shown that the portal is much more like a sensor that essentially helps measure out an appropriate length of DNA for each capsid particle, ensuring faithful production of new viral particles.”
The finding solves a longstanding puzzle in the field, and reveals a potential drug target for one of the most common human viral pathogens, herpesviruses, which is responsible for diseases such as chicken pox, mononucleosis, lymphomas and Kaposi sarcoma.
Cingolani and colleagues started their work 18 years ago by characterizing the structure of the portal protein using X-ray crystallography from P22, a bacteria-infecting virus that has a nearly identical portal protein as the human-infecting herpesviruses. In a paper published in 2011, the group showed that the portal protein looked like a perfect doughnut on a pedestal, with 12-fold rotational symmetry. Paradoxically, though, the protein wasn’t very good at binding to DNA, which should have been an essential ability, the researchers thought.
“We figured the portal protein we had studied for over a decade must be an end-stage, or mature version, of a more plastic and dynamic molecular machine,” says Cingolani. “And that it must also adopt other conformations earlier in viral assembly that have the capacity to bind both DNA and other motor proteins, or terminases.”
Proteins can change structure and hence their function and biochemical activity many times before reaching their final mature state. While these intermediate states are unstable and sometimes exist for tiny fractions of a second, they can also have important functions.
In the current paper, Cingolani and colleagues describe their success in identifying and characterizing an immature state of the portal protein, of which they determined the three-dimensional structure down to the atomic level. Unlike the mature final-stage protein that was profoundly symmetric, this immature conformation of portal protein is surprisingly asymmetric and has the ability to bind strongly to both the motor and the DNA itself.
“We think that DNA binds to the immature portal protein and wraps around it like a python, as it enters the viral capsid with the help of the motor protein. This DNA stranglehold causes the portal protein to begin to transform into its final symmetric state that because of its weak binding will ultimately release both the DNA and the motor, cutting off the DNA-loading at an appropriate length,” says Cingolani. “It’s a completely novel mechanism for sensing DNA. It’s a conformational change from asymmetric to symmetric that’s completely unexpected, yet makes perfect sense.”
In addition, the portal protein is unique to viruses, which makes it — in all of its various forms — a potentially good drug target. Because some herpesviruses infect and lay dormant in human cells until they reawaken by stress, developing a therapy that could interfere with viral production at different levels could prove a useful therapeutic strategy.
“It took us 18 years to understand that the portal protein functions by existing in two states that turn the viral DNA packaging on and off by changing its structure. At 18, it feels like this story has come of age along with the research,” says Cingolani.
This work was supported by NIH grant R01 GM100888 and GM076661, by the AAAS Marion Milligan Mason Award for Women in the Chemical Sciences. The Sidney Kimmel Cancer Center X-ray Crystallography and Molecular Interaction Facility is supported in part by NCI grants P30 CA56036 and S10OD017987.
Source: Thomas Jefferson University




