Researchers from the University of California San Diego developed a way to minimize bystander edits, unwanted edits that occur when a base editor edits the bases surrounding the desired nucleobase. The work is published in Nature Biotechnology.
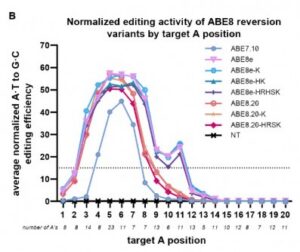
ABE 8 variants show a higher editing efficiency, but a larger editing window. ABE7.10 shows the opposite: a smaller window, but lower efficiency. Credit: Alexis Komor lab / UC San Diego
Base editing has progressed from the laboratory to patient care, treating debilitating or terminal illnesses, in less than a decade. A type of gene editing that makes chemical changes to our DNA, base editing was developed by Alexis Komor, associate professor in the Department of Chemistry and Biochemistry at the University of California San Diego.
The bystander effect
Base editing makes an edit to a single strand of DNA. In the case of adenine base editors (ABEs), a target adenine (A) nucleobase can be edited to guanine. However, if multiple A bases are neighboring the target, sometimes all the A bases are edited, even if there was only one A target. These bystander edits may be harmless or “silent,” or they may bring about unwanted consequences, including cell death.
One way to reduce bystander editing is to narrow the “editing window,” or the stretch of bases the ABE can effectively edit. However, research has shown that narrowing the editing window usually lowers the efficiency of desired on-target editing as well. The researchers wanted to develop a way to lower the incidence of bystander editing while maintaining the efficiency of on-target editing.
“Especially for therapeutic applications, it’s important to optimize both properties. A genome editor could be quite efficient at installing a desired edit, but if it does so concurrently with other, unwanted edits, then it won’t be safe to use as a therapeutic,” Komor said.
For this work, Mallory Evanoff, a chemistry alumna and former postdoctoral scholar in Komor’s lab, focused attention on an initial version of ABE called ABE7.10. ABE7.10 has a narrower editing window than more recently engineered ABE8.20 and ABE8e editors, but also has a lower editing efficiency. ABE7.10 edits positions 4 to 8, ABE8.20 expands to 3 to 9 and ABE8e expands to 3 to 11. The ABE 8 variants have higher on-target editing and, thus, are more widely used (including in clinical trials), but they also have much higher bystander editing.
To convert A bases to G bases, ABE7.10 uses an engineered enzyme with 14 point mutations installed that were previously identified to be important for effective base editing. The process that introduced and selected for these important mutations (a technique called directed evolution that won the Nobel Prize in Chemistry in 2018) was completed in E. coli bacteria cells — a different environment from the human cells where base editors are ultimately applied. This mismatch meant some mutations that were beneficial in bacteria turned out to be unnecessary or even counterproductive in human cells, and it was unclear how each of the 14 mutations contributed to editing activity.
To untangle the effects, the researchers used reversion analysis, reverting each mutation back to its original wild-type sequence. They characterized the activity of all 14 of the individually reverted mutations in both human cells and E. coli bacterial cells.
The researchers noticed that some of the mutations behaved similarly in bacteria, while others had disparate effects in the two host systems. Given that most researchers’ usage of ABEs is in human cells, they identified five mutations that, when reverted individually, either had no impact or increased editing activity in human cells. The scientists then combined the five mutations to create what they call a minimally evolved ABE, or ME-ABE.
The resulting ME-ABE keeps a narrow editing window, but gains editing activity similar to the two ABE 8 variants that are more commonly used, matching the activity of ABE8e and ABE8.20 at the central editing position (position 5) in most sequence contexts.
“This is one of the first times that we have been able to decouple the characteristics of efficiency and editing window. In other work, increasing editing activity has always come with the cost of a wider editing window, and thus more bystander editing,” stated Komor. “We’re excited to offer this tool to the community to help researchers who want to model or correct mutations and have unacceptable bystander bases next to their target A’s”.
Cracking the code
“The thing about building these precision tools is that we can install those mutations in model organisms and ask the right questions before we try any therapies in patients,” stated Mallory Evanoff, one of the researchers. “Or we can tease apart, very specifically, the fact that many people might have a whole host of mutations, but only a handful of them are contributing to whatever symptom or disease we’re working on.”
Evanoff calls the ME-ABEs “cut-both-ways” tools because they help researchers better model and understand possible genetic mutations, and let them implement potential corrections and build personalized medicines that can cure specific diseases. In the study, ME-ABEs were used to successfully edit six single-nucleotide variants of clinical interest that had previously proved too difficult to edit precisely with existing tools.
While Komor and Evanoff work on developing new base editors — particularly by evolving them in mammalian cells instead of bacteria — they also hope ME-ABEs are a jumping off point for other researchers seeking to develop effective gene-editing tools.
“One of the great things about working in the tool development world is that you’re making something that allows other people to answer their own questions,” stated Evanoff. “These tools are going to be available for researchers who are really, really well-informed on their particular disease or their particular system and now have an ability to model a cure.”
Plasmids for this and other work from Komor’s lab can be found on AddGene (https://www.addgene.org/Alexis_Komor/).