by University of California San Diego
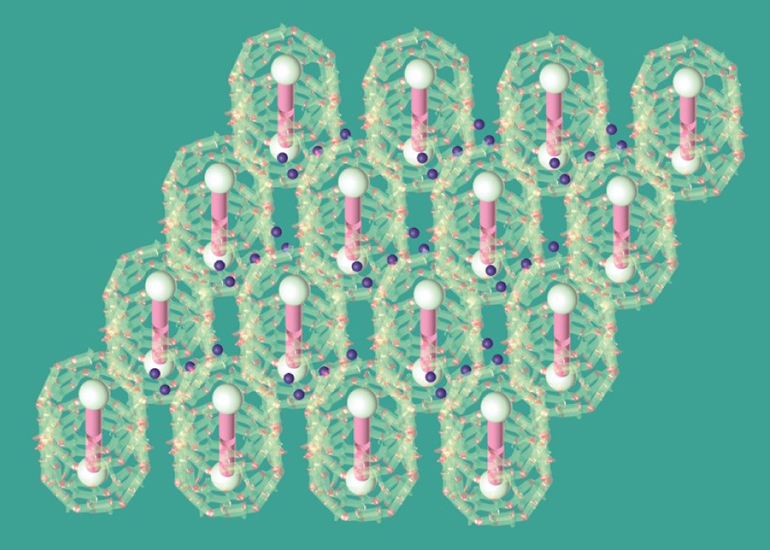
Crystal structure of the self-assembled materials. Blue dots represent strongly bound water. The variation of the number of water molecules reflects the potential heterogeneity of the strongly bound water numbers in each site. Credit: Haoyuan Wang, UC San Diego
As a magnifying lens is to a detective, a microscope is to a chemist. These tools of trade help both investigator-types follow the evidence. In the case of a detective, it’s to find the marks of a criminal; in the case of a chemist — at least for UC San Diego’s Wei Xiong — it’s to discover molecular properties that influence chemical reactions and materials’ functions.
When it comes to gathering clues, Xiong hunts down miniscule molecules that aren’t easy to see, which is why he and his team of researchers developed a microscope that gives them a thorough view of molecular systems — not just single traits of molecules. Their project’s results are now published in an article in Proceedings of the National Academy of Sciences.
He and members of the Xiong Group used their new instrument, called a transient vibrational sum-frequency generation microscope, to inspect hydrogen bond (H-bond) interactions, which are critical for self-assembled soft materials — an important area of research for synthetic biology/biomimetics. The delicate H-bond interactions balance electrostatic interactions needed for synthetic lattice self-assemblies to resemble the form and function of their biological analogs (e.g., cells).
The novel tool is devised to study molecular interfaces and self-assembled materials spatially, temporally—meaning their ultrafast dynamics—and their energy. So the researchers, who included lead author Haoyuan Wang, a fifth-year graduate student who co-designed the experiment, applied it to the study of recently developed self-assembled materials that mimic biological entities for biomedical purposes. According to Xiong, although these materials have been developed, the physical driving force that can make them mimic the crystallinity and flexibility of biological organisms, such as viruses, remains unclear.
The microscope helped the scientists see that different self-assemblies can have distinct H-bond interactions, depending on their levels of hydration. Additionally, the scientists found that the H-bond interactions are ordered in the self-assemblies, i.e., the H-bond only exists with certain molecular groups, while other groups nearby don’t have it.
“On the other hand, the H-bond can break and restore fast — on the hundreds of the femtosecond time scale,” said Xiong. “We believe the local ordering and fast dynamics of H-bond is the key for this material to mimic biological crystallinity and flexibility, and thereby the local hydration of each self-assembly set determines the unique H-bond interactions and its distinct mechanical properties. This fundamental knowledge offers guidelines to further develop biomimetic materials for biomedical applications.”
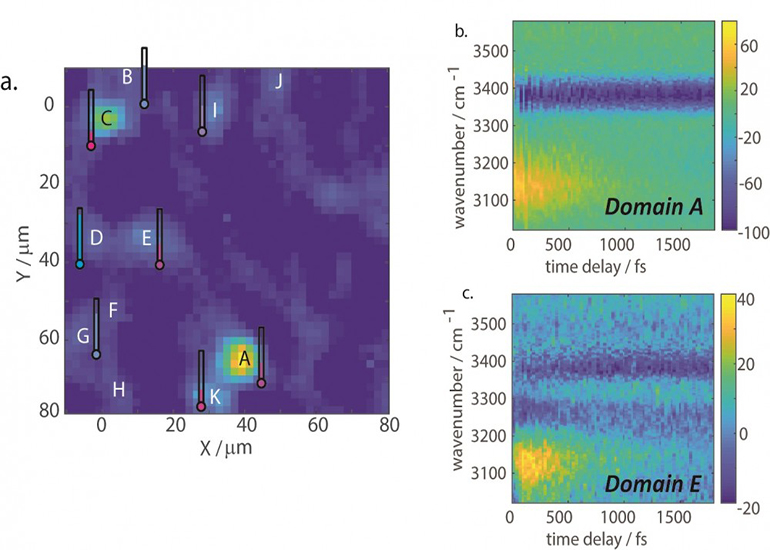
a)VSFG intensity image (PPP polarization) for the OH region of molecular self-assembled micron sheets. The hygrometer next to each domain represents the relative hydration level. (b and c) Two representative types of vibrational dynamics of different domains. Credit: Haoyuan Wang, UC San Diego
Xiong said that one hypothesis is that H-bond interaction between water and the materials is essential. Since the H-bond interaction in these materials is ultrafast (at the femtosecond to picosecond scale), multiple interactions’ dependency on the morphology, or form, of the self-assemblies has not been studied much, he noted. Therefore, the team’s new microscopy technique is positioned to resolve this question from multiple aspects.
“The development and demonstration of this technique make it available for other chemical systems, such as aerosol surfaces, which are related to environment and public health,” noted Xiong.
The professor of article admitted that the project was challenging since there were few previous examples of how to develop a microscope that not only captures an image but also reveals the energy level and ultrafast dynamics of the systems. He said that the technique was developed through hard work and creativity, which resulted in the collection of a massive amount of 4D hypercube data (2D in space, 1D in time and 1D in energy) that posed the challenge of extracting useful information from it —“like a detective finding the key clue from 10,000 threads.”
Xiong said they overcame this difficulty by first analyzing the data on a coarse level and then selecting the interesting data point to perform detailed analysis. In the future, he expects to combine it with artificial intelligence (AI).
“Because we applied a new tool on new material, distilling the key and new molecular physics was the final challenge,” said Xiong. “This step occurred during COVID, and Haoyuan and I spent a lot of time on Zoom and exchanged emails until we developed a simple and intuitive model to convert the H-bond dynamics into local hydration level, demonstrating the critical new insights we can learn using this technique.”
Wang said that during the study they struggled with a technical problem, which, at one point, stopped them from making progress. Xiong noted that Wang had just joined the group when the project began, so he had little knowledge of the instrument. Still, he introduced the unique self-assembled system, which is the core system the group studied.
“I still remember it was right before Christmas, and I was on my way to join the department party,” said the PhD student. “But when I was halfway, I realized a method to resolve the problem. I shouted to myself and ran back to the lab to test my thought immediately. Though I missed the department party, I enjoyed that moment when I got inspired.”
Xiong said that in figuring out every detail of the project over several years, he became a more experienced mentor, while he enjoyed seeing his students grow through the process. “Haoyuan became an expert in nonlinear optics who came up with many brilliant ideas to tackle the technical and scientific difficulties faced during this project,” said Xiong. “I was surprised to learn how ordered the H-bond interaction was on a mesoscopic scale: the interaction and local hydration are uniform in each self assembly, but differ between self assemblies. Before this work, my perception — and the general perception — was that the H-bond network is homogeneous. Clearly, these works show that self-assembly structures influence H-bond interaction.”
Other members of the Xiong Group who contributed to this research included Jackson Wagner, Wenfan Chen and Chenglai Wang. This study was supported by DARPA (D15AP00107); DOE, BES (DE-SC0019333) and NSF (CHE-1801971, CHE-1808111 and CHE-1828666).




Tell Us What You Think!