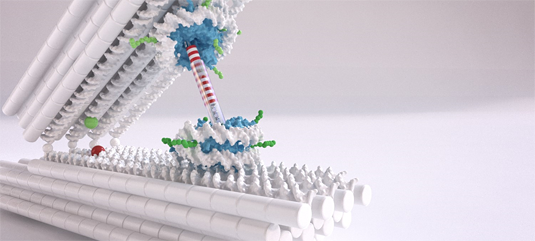
Image: Chris Hohmann (NIM); Dietz Lab (TUM)
The mode of packaging of the genomic DNA in the cell nucleus determines patterns of gene expression. Munich researchers have used DNA-based nano-tweezers to measure the forces between nucleosomes, the basic packing units of nuclear DNA.
Every human cell contains some two meters of deoxyribonucleic acid (DNA), which encodes the genetic information that specifies the cellular structures and functions. Moreover, this “genomic” DNA is packed into the cell nucleus, which is less than 10 micrometers in diameter. This means that the nuclear DNA must be packed, primarily by interacting with specific proteins. The basic packing unit is a particle made of proteins called histones, around which the DNA is wrapped. Comparable to small spools, these structures are referred to as nucleosomes. Nucleosomes in turn are linked to one another by segments of DNA that extend between the core particles and are not wrapped around them. Viewed under the electron microscope, the DNA packed in nucleosomes resembles beads on a string.
The next level of packaging involves the mutual interaction of nucleosomes, and the resulting higher-order structures have not yet been completely characterized. A team of scientists led by Hendrik Dietz of the Technical University of Munich and Philipp Korber at LMU’s Biomedical Center has now taken a substantial step toward solving this puzzle: For the first time ever, they have succeeded in directly measuring the attractive forces that act between nucleosomes. Their results appear in the journals Science Advances and Nano Letters.
Dietz, holder of the Chair for Experimental Biophysics at the TUM, uses DNA as a construction material to build molecular structures — a technology referred to as DNA origami. He and his team have now used the method to create structures consisting of two rigid DNA bars connected by a flexible joint that acts as a spring. These can be used like tweezers to measure the strength of the interactions between nucleosomes. One nucleosome is attached to each arm of the tweezers.
“We can control the position and orientation of the nucleosomes in the DNA tweezers with a very high degree of precision,” says Dietz. “This is very important when it comes to really being able to measure the interactions.”
The LMU researchers took on the task of developing nucleosome structures that can be integrated into the tweezers. Philipp Korber, Privatdozent and Group Leader at the Chair of Molecular Biology at the BMC, explains: “Normally the two double-stranded ends of the DNA spooled around the nucleosome are very close to one another. But what we needed were two protruding single strands, closer to the middle. This was a significant issue, since such a configuration can destabilize the entire structure. Our team member Corinna Lieleg nevertheless succeeded in finding the right spots for these handles.”
The researchers were able to measure a very weak interaction between integrated nucleosomes, equivalent to an attractive force of 1.6 kcal/mol, at a range of about 6 nanometers (nm). The orientations of the nucleosomes relative to one another were found to have hardly any effect. However, particular chemical modifications in the histone proteins further weakened the interactions.
The result could help resolve a current scientific dispute. According to the current theory, nucleosomes form a type of super-spiral with a diameter of 30 nanometers, the so called 30-nm fiber. So far, however, this higher-order 30-nm structure has never been observed in living cells. Whether or not the chromatin really takes on the form of such a super-spiral is still highly controversial. Indeed, the minute attractive forces between the nucleosomes, which the researchers have now successfully measured, appear to contradict the theory.
“Our data point to very soft structures that are easily deformed by external influences,” says Dietz.
How nucleosomes are organized into higher-order structures is a fundamentally important issue, as it has profound implications for the control of gene expression. Only those genes which lie within relatively non-compact chromatin are accessible to “activation,” which allows the proteins they encode to be produced by the cellular machinery.
“Over the past ten years it has become clear that many of the changes and mutations that transform cells into cancer cells take place at this level,” Korber says. In a cancer cell, the normal mechanisms that determine which genes are active and which are inactive are disrupted. Genomic regions which should not be accessible are left open and vice versa. “However, if only the packaging is defective, and not the gene itself, it should in principle be possible to restore the proper packaging again.”
The researchers plan to use the molecular tweezer technique to investigate other structures. “In biology the orientation of structures with respect to one another is always important,” says Korber. “Now we have a kind of molecular clamp which we can use to specifically control the spatial orientation of structures to one another.”
In another experiment, the researchers had measured the force needed to unwind the DNA from the nucleosome. Thus, the DNA-based origami tweezers can be used to measure the forces acting between and within nucleosomes.




