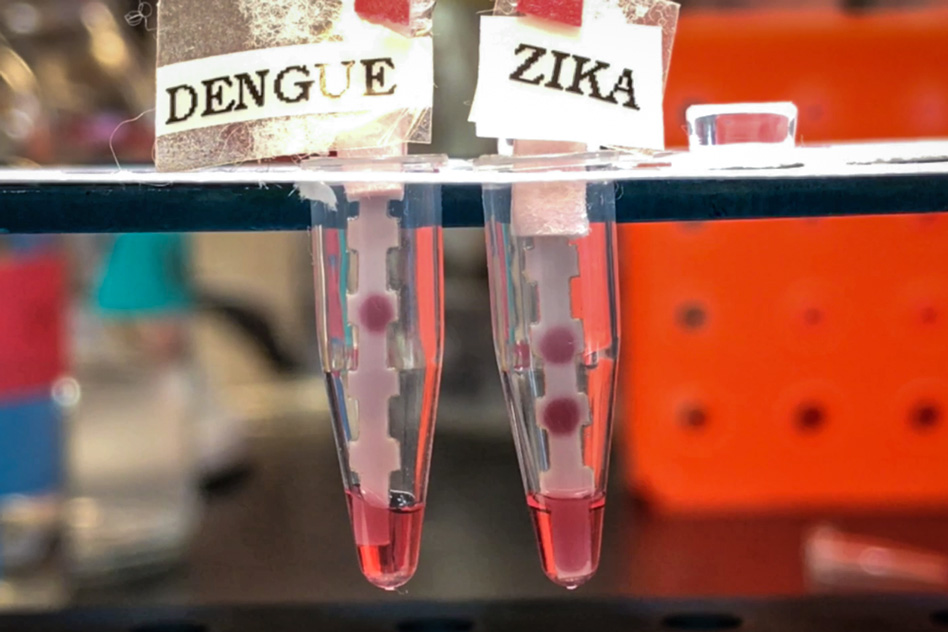
“It’s important to have a single test that can differentiate between the four serotypes of Dengue and Zika, because they co-circulate. They’re spread by the same mosquito,” says Kimberly Hamad-Schifferli, an associate professor of engineering at the University of Massachusetts at Boston and a visiting scientist in MIT’s Department of Mechanical Engineering. Courtesy of the researchers
MIT researchers have developed a paper-based test that can diagnose Zika infection within 20 minutes. Unlike existing tests, the new diagnostic does not cross-react with Dengue virus, a close relative of the Zika virus that can produce false positives on many Zika tests.
This test could offer an easy-to-use, cheap, and portable diagnostic in countries where Zika and Dengue are both prevalent and the gold-standard test that measures viral RNA in the bloodstream is not available.
“It’s important to have a single test that can differentiate between the four serotypes of Dengue and Zika, because they co-circulate. They’re spread by the same mosquito,” says Kimberly Hamad-Schifferli, an associate professor of engineering at the University of Massachusetts at Boston, a visiting scientist in MIT’s Department of Mechanical Engineering, and a co-senior author of the paper.
The researchers worked with scientists around the world to test the new device on patient samples and confirmed that it can accurately distinguish Zika virus from related viruses.
Lee Gehrke, the Hermann L.F. von Helmholtz Professor in MIT’s Institute for Medical Engineering and Science (IMES), is also a senior author of the study, which appears in the Sept. 27 issue of Science Translational Medicine. The paper’s first authors are IMES research scientist Irene Bosch and Department of Mechanical Engineering postdoc Helena de Puig.
No more false positives
One of the biggest challenges in diagnosing Zika is that many of the tests are based on antibodies that interact with a viral protein called NS1, which is found in the bloodstream of infected patients. Unfortunately, many other viruses from the same family, known as flaviviruses, have similar versions of NS1 and can produce a false positive. Flaviviruses include West Nile virus and the virus that causes yellow fever, as well as Dengue virus.
In an effort to create a more precise diagnostic, the MIT team set out to find antibodies that would interact exclusively with NS1 protein produced by the Zika virus, as well as antibodies specific to NS1 from each of the four different strains of the Dengue virus.
To achieve this, the researchers exposed mice to Zika and Dengue viruses and then screened the resulting antibodies, in pairs, against every flavivirus’ version of the NS1 protein. This allowed them to identify pairs of antibodies that react only with one version of NS1 and not any of the others.
“We knew by informatics analysis that if we looked enough, and we teased out the repertoire of the B cells of these animals, we would eventually find those antibodies,” Bosch says. “We were able to tease out the very few antibodies within the repertoire that would give you uniqueness in the detection.”
The researchers used these pairs to create five separate tests, one for each virus. They coated strips of paper with one antibody from each pair, while the second antibody was attached to gold nanoparticles. After adding the patient’s blood sample to a solution of these nanoparticles, the paper strip is dipped into the solution. If the target NS1 protein is present, it attaches to the antibodies on the paper strip as well as the nanoparticle-bound antibodies, and a colored spot appears on the strip within 20 minutes.
This approach requires five test strips per sample to test for each virus, but the researchers are now working on a version that would test for all five with one strip.
Most countries where Zika and Dengue are prevalent do not allow patient samples to be shipped out of the country, so the researchers traveled to several countries, including Mexico, Colombia, India, and Brazil, to test their devices with patient samples.
They found that their results were comparable to those obtained by polymerase chain reaction (PCR) tests, which detect viral RNA in the bloodstream. PCR tests are not widely used in areas where Zika virus is found because they require trained personnel and lab equipment that are not available everywhere.
“Since conventional methods require a great deal of time for sample collection and diagnostics, this inexpensive, paper-based, rapid diagnostic will be very useful for the diagnosis of many infectious diseases,” says Luke Lee, an associate president of the National University of Singapore and director of the Biomedical Institute for Global Health Research and Technology in Singapore.
Emerging viruses
Dengue infects hundreds of millions of people annually, mostly in tropical regions. It is usually not fatal, but in areas where there is more than one serotype circulating, it is more likely to produce a severe, potentially life-threatening illness. A diagnostic that can distinguish between all four serotypes of Dengue fever could give doctors a way to discover early on when a new serotype has entered their region.
“When we have traveled to the places where these viruses are problems, the people there unanimously say that they need more surveillance. They need to know which viruses are circulating in their environments,” Gehrke says.
The researchers believe that their approach should also enable them to quickly develop diagnostic tests for other related viruses that might emerge in the future.
“By already screening this group of antibodies that we have against all these antigens we have, like West Nile, we already know how well they react. So that’s information we could use in the future to develop additional tests that can be used to detect other emerging viruses,” Gehrke says.
They are now working on a diagnostic for the emerging Powassan virus, which is carried by the same tick that spreads Lyme disease. Powassan, found mainly in the northeastern United States and the Great Lakes region, causes a severe form of encephalitis.
The research was funded by the U.S. Public Health Service and the Science, Technology and Innovation Fund of Colombia.