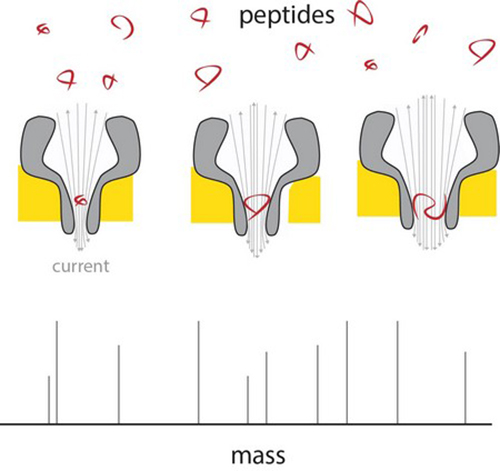
A peptide enters the thin end of the nanopore, and there changes the current in proportion to its mass. By using differently sized nanopores, a range of peptide sizes can be measured. | Illustration G. Maglia
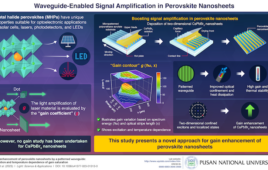
Scientists of the University of Groningen have developed nanopores that can be used to directly measure the mass of peptides. Although the resolution needs to be improved, this proof of principle shows that a cheap and portable peptide mass spectrometer can be constructed using existing nanopore technology and the patented pores that were developed in the lab of UG Associate Professor of Chemical Biology, Giovanni Maglia.
An article on this discovery was published in Nature Communications.
Mass spectrometers are invaluable for studying proteins, but they are both bulky and expensive, which limits their use to specialized laboratories.
“Yet, the next revolution in biomedical studies will be in proteomics, the large-scale analysis of proteins that are expressed in different cell types,” says Maglia.
For although each cell in your body carries the same DNA, the production of proteins differs hugely between cell types. “And also, proteins are modified after they have been produced, for example by adding sugars that can affect their function.”
Nanopore technology could offer a way to analyses single molecules. In previous work, Maglia already showed that biological nanopores can be used to measure metabolites and to identify proteins and peptides. These pores are large protein structures, incorporated in a membrane. Molecules entering a pore or passing through it cause a change in an electric current across the pore.
“A problem in measuring the mass of peptides is that they pass too quickly through even the smallest
Making smaller pores was a challenge. “Pores are made up from a number of monomers, so we initially modified the interaction between these monomers, but that didn’t work.”
The observation that mixing monomers with larger amounts of lipids—which make up the membrane—resulted in a larger percentage of smaller pores gave Maglia and his team the idea to modify the interaction between monomers and lipids. This indeed resulted in pores made up from a smaller number of monomers, which reduced pore size.
Maglia was then able to produce funnel-shaped pores which, at their narrow end, only measured 0.84 nanometers. “These are the smallest biological pores ever produced.”
The next challenge was to ensure that peptides would pass through the pores, irrespective of their chemical composition. “The pores have a negative charge, which is necessary for their proper function,” explains Maglia.
The charge causes water to flow through the pore, dragging the peptides along. But negatively charged peptides would be repelled by the negative charge at the thin funnel end. Maglia modified the charge by altering the acidity of the fluids used.
“Eventually, we managed to find the right conditions by setting the acidity at a pH of exactly 3.8. This allows negatively charged peptides to pass through, while maintaining a large enough water flow through the pores.”
Measurements across nanopores of different sizes show that the electric current is linear with the volume of the peptide passing through. These peptides ranged from 4 to 22 amino acids in length. The difference between the amino acids alanine and glutamate could be measured in this system, which meant the resolution is around 40 Dalton (a measure for protein mass).
“The resolution of conventional mass spectrometers is much better, but if we could get the system about forty times more sensitive, it would already be useful in proteomics research,” says Maglia.
There are a number of ways to improve the resolution, says Maglia. “We could engineer the nanopore with artificial amino acids, or use different ions in our solutions, reduce the noise by changing the temperature etcetera.”
The nanopore system has several unique selling points: it measures single molecules, the technology itself is already commercially available and it is relatively cheap. Furthermore, the nanopore system is portable. And, by using many different pores in a device, you can simultaneously measure differently-sized peptides and even peptide modifications.
“All of this means that a versatile and cheap mass spectrometer for peptide analysis is feasible,” says Maglia. “And that would mean that more laboratories would be able to afford to conduct very important proteomics studies.”